Cost of clc genomics workbench5/6/2023  
Summerer D (2009) Enabling technologies of genomic-scale sequence enrichment for targeted high-throughput sequencing. Ross MG, Russ C, Costello M, Hollinger A, Lennon NJ, Hegarty R, Nusbaum C, Jaffe DB (2013) Characterizing and measuring bias in sequence data. Reich D, Green RE, Kircher M, Krause J, Patterson N, Durand EY, Viola B, Briggs AW, Stenzel U, Johnson PL, Maricic T, Good JM, Marques-Bonet T, Alkan C, Fu Q, Mallick S, Li H, Meyer M, Eichler EE, Stoneking M, Richards M, Talamo S, Shunkov MV, Derevianko AP, Hublin JJ, Kelso J, Slatkin M, Paabo S (2010) Genetic history of an archaic hominin group from Denisova Cave in Siberia. Poortvliet M, Hoarau G (2013) The complete mitochondrial genome of the Spinetail Devilray, Mobula japanica. Mason VC, Li G, Helgen KM, Murphy WJ (2011) Efficient cross-species capture hybridization and next-generation sequencing of mitochondrial genomes from noninvasively sampled museum specimens. J Fish Biol 80:1075–1119ĭulvy NK, Fowler SL, Musick JA, Cavanagh RD, Kyne PM, Harrison LR, Carlson JK, Davidson LN, Fordham SV, Francis MP, Pollock CM, Simpfendorfer CA, Burgess GH, Carpenter KE, Compagno LJ, Ebert DA, Gibson C, Heupel MR, Livingstone SR, Sanciangco JC, Stevens JD, Valenti S, White WT (2014) Extinction risk and conservation of the world’s sharks and rays. Furthermore, given the low cost of NGS, the percentage of reads on target should be adequate for most research.Ĭouturier LI, Marshall AD, Jaine FR, Kashiwagi T, Pierce SJ, Townsend KA, Weeks SJ, Bennett MB, Richardson AJ (2012) Biology, ecology and conservation of the Mobulidae. However, the efficiency of hybridization will be taxa specific, depending on the phylogenetic distance between the taxa use to design the baits and the target but also on features of the mitogenomes such as GC contents and gene rearrangements. Optimization of the hybridization step could results in improved enrichment efficiency, and increased coverage. The percentage of mtDNA read was 1–2 order of magnitude higher than what was obtained by direct sequencing without targeted enrichment. Although the percentage of on-target reads was overal quite low (1.8–19.2 %) the sequencing run resulted in adequate coverage (average coverage depth = 9–833 reads) of large part of the mitogenomes (coverage = 84–99 %) of the 11 species (Table 1). Sequencing of the remaining two degraded samples produced no barcoded reads. Most of the remaining 15.500 bp of the mitogenome (84 % or more) was successfully sequenced for 11 of the 13 species, including two out of four heavily degraded samples (for which PCR amplifications of short mtDNA fragments were unsuccessful). 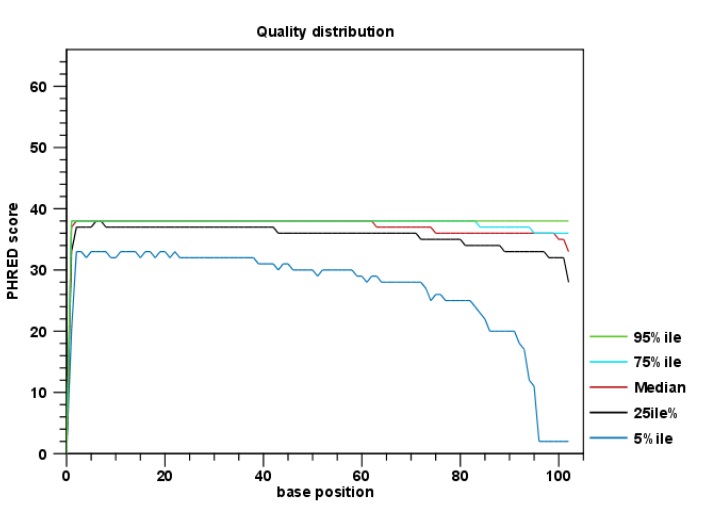
The mitochondrial Control Region and flanking regions were covered by very few reads in all species, due to the presence of long AT-rich tandem repeat regions (Poortvliet and Hoarau 2013), which are known to lead to coverage bias on the Ion Torrent sequencing platform (Ross et al. japanica in CLC Genomics Workbench v.6 (CLC bio, Denmark) using default mapping parameters (Mismatch cost 2, insertion cost 3, deletion cost 3, length fraction 0.5, similarity fraction 0.8). Sequence reads belonging to each barcoded library were mapped to the reference genome of M. The enriched amplified library pool was purified using the PureLink PCR Purification Kit (Life Technologies), quantified and diluted appropriately. Post capture amplification for ten cycles was performed using the Library Amplification Primer Mix supplied in the Ion Plus Fragment Library Adapters Kit (Life Technologies). Recovery of the captured targets with MyOne Streptavidin C1 magnetic beads (Life Technologies) was followed by elution and cleanup of the enriched library pool. 
Hybridization was performed for 36 h at 65 ☌. Two custom oligonucleotide blocking-probes were used to prevent the cross-hybridization between Ion Torrent adapters during the hybridization step (BlockingAbc: ATCIIIIIIIIIICTGAGTCGGAGACACGCAGGGATGAGATGG 3′-PHO and BlockingP1: ATCACCGACTGCCCATAGAGAGGAAAGCGGAGGCGTAGTGG). 
As reference sequence for the current study we used the Mobula japanica complete mitochondrial genome (NC_018784), which is 18.880 bp long (Poortvliet and Hoarau 2013). More specifically, we used the Mybaits1 system, which contains a custom library of 20.000 biotinylated 120mer single stranded DNA baits designed against a specific reference sequence. Sequence enrichment for targeted sequencing was achieved using a customizable liquid-phase DNA capture system under the commercial name MYbaits (). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
